Multivoxel Pattern Analysis (MVPA) is the latest big thing in the neuroimaging world. MVPA is a multivariate statistical technique that can be applied to fMRI brain scan results as an alternative to conventional univariate methods of finding brain activation. Neuroscientists love MVPA for two reasons: first, it offers more 'blobs for your buck' - it often detects neural signals that don't show up on conventional scans. Secondly, it seems to offer a way to go beyond merely detecting and localizing activity and actually provide insights into how information is represented in the brain. MVPA promises to not just locate but also 'decode' the brain's activity, one of the Holy Grails of modern neuroscience. But Texas-based neuroscientists Tyler Davis and colleagues offer an important note of caution in a new NeuroImage paper: What do differences between multi-voxel and univariate analysis mean? They show convincingly that we can't assume that MVPA tells us anything special about the neural code.
Because univariate voxel-wise tests and MVPA differ in their ability to detect multidimensional effects, it is tempting to conclude that MVPA tests have identified a multidimensional code for a variable when MVPA results are significant but voxel-wise tests are not. [ But ] the validity of this inference depends upon how likely differences between univariate voxel-wise analysis and MVPA are to arise when only a single dimension underlies activation patterns.
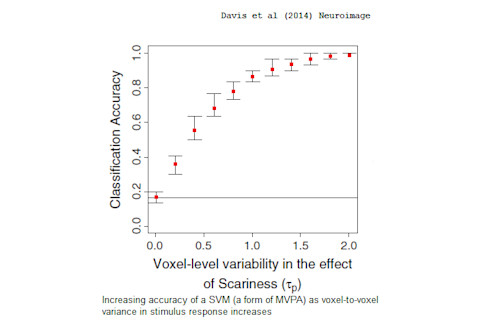
In other words, if MVPA could outperform conventional methods, even in the absence of any kind of sexy neural code waiting to be decoded, then we shouldn't read too much 'decoding' into those results. Using simulations, Davis et al confirm that, indeed, we don't need to assume any kind of complex neural encoding to explain why MVPA is often more sensitive than other methods. Rather, the superiority of MVPA could arise from a very simple source - differences in the brain response to a stimulus across different regions of the brain:
Our results indicate that MVPA is sensitive to voxel-by-voxel variability in the parameters (beta weights) relating activation within voxels to experimental variables, even when these voxel-wise parameters are draws from the same unidimensional distribution […] In cases in which an underlying neural space is unidimensional but high subject-level variability renders voxel-wise tests non-significant, many common MVPA tests neutralize this subject-level variability and will be significant as long as there is reliable voxel-level variability within subjects.
As they point out, the existence of voxel-by-voxel variability in stimulus response is a fundamental assumption of neuroimaging (otherwise, the brain would be one homogenous lump). This variability exists in every single fMRI experiment. Yet that variability is all we need to explain the success of MVPA.
MVPA may only indicate what is already assumed in most contexts in which the voxel-wise analysis is employed - that the effect of an experimental variable varies across voxels... [it] does not necessarily allow for any special conclusions about how information in the underlying activation patterns is encoded, or [indeed] that an effect differs in any other substantive way from a simple linear mapping of a single experimental variable onto voxels within a region.
Davis et al do hold out some hope, however, saying that we can indeed study neural codes using fMRI but that it will require more sophisticated methods. For example, if we explicitly model a hypothetical neural encoding scheme, we could use fMRI to test that hypothesis. We just can't throw MVPA at some data, with no hypothesis in mind, and expect it to crack the code for us. A year ago I blogged about another paper that poured cold water on the over-interpretation of MVPA. The issue last time around was that MVPA may detect areas where activation differs across individuals, even if those differences are unrelated to neural coding. Davis et al are making a separate (and complementary), point which is that MVPA is also sensitive to spatial differences in the neural response to a stimulus. The story in both cases is that MVPA is so powerful that it can exploit (what most neuroscientists would call) trivial differences.

Davis, T., LaRocque, K., Mumford, J., Norman, K., Wagner, A., & Poldrack, R. (2014). What do differences between multi-voxel and univariate analysis mean? How subject-, voxel-, and trial-level variance impact fMRI analysis NeuroImage, 97, 271-283 DOI: 10.1016/j.neuroimage.2014.04.037