As many of you know around the year 2000 the analyses of Y chromosomal human lineages became a pretty big deal. The reason these lineages are important and useful is that they record the uninterrupted ancestry of males, from father to son, along the Y chromosome. Instead of the complexities of the whole genome, as with mtDNA you have a simple and elegant phylogenetic tree to interpret. The clusters along this tree are defined as broad haplogroups, united by derived states from a common ancestor. One of the largest haplogroups is R1a1a. It happens to be my paternal lineage, as well as Dr. Daniel MacArthur’s and Dr. Zack Ajmal’s.
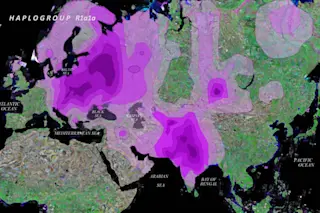
The map above illustrates the peculiarity of R1a1a: it is geographically enormously expansive. How to explain this distribution? A naive response might be that this distribution is surprising similar to that of the Indo-European languages. Unfortunately this runs up against the conundrum that low caste South Indian groups, relatively untouched by Indo-Aryan culture (at least until the past few hundred years), also manifest high frequencies of R1a1a.
To make a long story short it seems that R1a1a is an old haplogroup with a lot of structure across Eurasia. Maju points me to a paper in American Journal of Physical Anthropology which simply & elegantly brings home to us some obvious insights, New Y-chromosome binary markers improve phylogenetic resolution within haplogroup R1a1:
Haplogroup R1a1-M198 is a major clade of Y chromosomal haplogroups which is distributed all across Eurasia. To this date, many efforts have been made to identify large SNP-based subgroups and migration patterns of this haplogroup. The origin and spread of R1a1 chromosomes in Eurasia has, however, remained unknown due to the lack of downstream SNPs within the R1a1 haplogroup. Since the discovery of R1a1-M458, this is the first scientific attempt to divide haplogroup R1a1-M198 into multiple SNP-based sub-haplogroups. We have genotyped 217 R1a1-M198 samples from seven different population groups at M458, as well as the Z280 and Z93 SNPs recently identified from the “1000 Genomes Project”.
The two additional binary markers present an effective tool because now more than 98% of the samples analyzed assign to one of the three sub-haplogroups. R1a1-M458 and R1a1-Z280 were typical for the Hungarian population groups, whereas R1a1-Z93 was typical for Malaysian Indians and the Hungarian Roma. Inner and Central Asia is an overlap zone for the R1a1-Z280 and R1a1-Z93 lineages. This pattern implies that an early differentiation zone of R1a1-M198 conceivably occurred somewhere within the Eurasian Steppes or the Middle East and Caucasus region as they lie between South Asia and Eastern Europe. The detection of the Z93 paternal genetic imprint in the Hungarian Roma gene pool is consistent with South Asian ancestry and amends the view that H1a-M82 is their only discernible paternal lineage of Indian heritage.
The table to the left shows you an Indian population from Malaysia. Malaysian Indians tend to be Tamils, from the south of the subcontinent. If they were finding individuals who were carriers of R1a1a, the data set is probably somewhat enriched for Tamil Brahmins and people of North Indian ancestry, though this does not alter the basic story. What you see is that all the Indians carry this one distinctive mutation. I find it unlikely that all these Malaysian Indians are Brahmins or North Indians, especially given that there is a non-trivial proportion of R1a1a in Tamil lower castes. So here you have a population with is probably representative of Indian Y chromosomal phylogeography before the Indo-Aryans arrived. Second, you see that M458 is well represented among Hungarians. This makes sense, insofar as this is a very common variant in Eastern Europe. Z280 also seems to be found in northern Eurasia. An interesting aspect is that in the Uzbek sample z93 has a high frequency. The Uzbeks are an admixed population. A Turkic component overlain atop an Iranian substrate. The frequency of Z93 suggests to me that the Eastern Iranians share common ancestry with South Asians. This is not a revolutionary finding, but it does imply that Z93 may have come, in part, with the Indo-Aryans (i.e., there were two or more waves of Z93).
The authors note that Z458 and Z93 carrying individuals exhibit “star like” phylogenies when STRs were analyzed. They are the top two panels. The Genghis Khan haplotype exhibits a star like phylogeny. In other words, it’s indicative of rapid expansion from a small founder group. In contrast, they argue that Z280 carrying Y chromosomes do not exhibit a star like phylogeny. The implication being that it did not undergo the same expansion. Dates of expansion (looking at the most recent common ancestor) for Z458 and Z93 are pegged to 7 and 10 thousand years before the present. I don’t put much stock in these dates personally, but I thought I’d relay them.
What can we say from this? If these results hold what they tell us is that R1a1a is a very lucky haplogroup, and its current range is a function of multiple expansions from a common and diverse R1a1a pool, probably in Central Eurasia. The presence of Z93 in Uzbeks, and Mongols, suggests to me that this variant was and is present in Iranians. Therefore, I don’t think that Z93 is indigenous to South Asia, but is intrusive. I believe it arrived with the “Ancestral North Indians.”