As a follow up to my post from yesterday, I decided to run TreeMix on a data set I happened to have had on hand (see Inference of Population Splits and Mixtures from Genome-Wide Allele Frequency Data for more on TreeMix). Basically I wanted to display a tree with, and without, gene flow. The technical details are straightforward. I LD pruned ~550,000 SNPs down to ~150,000. I ran TreeMix without and with migration parameters with the Bantu Kenya population being the root. Finally, when I did turn on the migration parameter I set it for 5. You can see the results below.
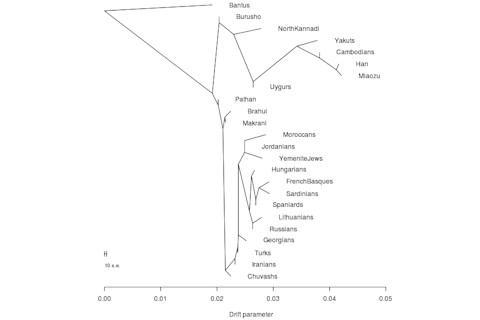
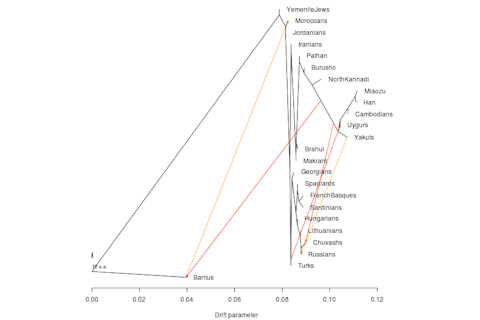
Most of the flows are pretty expected. The West Eurasian flow from the Turks to the Uygurs makes sense, because there is a large West Asian component to what the Uygurs have (from East Iranians?). The Chuvash are a Turkic group with minor, but significant, Turkic component. The HGDP Russian sample does have some East Eurasian ancestry. And the Moroccans also have African ancestry. But your guess is as good as mine with the Bantu flow in. These are I think Kenya, so it might be trying to interpret Nilotic admixture as generalized Eurasian. A minor note: installing TreeMix and generating the appropriate files from pedigree format is not to difficult. But you might have confusion in how to generate the pedigree input file. You do it like so in PLINK: ./plink --noweb --bfile YourFile --freq --within YourGroupNamesFile --out YourOutPutFile It's the last you want to put into TreeMix's python conversion script. The YourGroupNamesFile is basically the .fam file with an extra column, the population names for each individual.