High school senior Thomas Deerinck didn’t expect his life to change when he walked into his science class one day in the mid-1970s. Local research scientist Betty Barbour had come to share her stunning images of the microscopic world. She was hoping to recruit students for her electron microscopy training program, eager for new helping hands. Deerinck promptly enrolled.
Forty years later, his award-winning photos have appeared on the covers of scientific journals. He’s also one of the most senior researchers at the National Center for Microscopy and Imaging Research in La Jolla, California, where he and his colleagues expand the boundaries of what a microscope can see and do.
But as his subjects have gotten smaller over the years, the large streams of data the photos produce need heavy-duty processing.
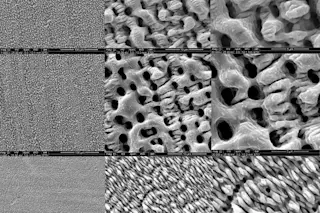
The jagged surface of a kidney stone. | Thomas Deerinck/NCMIR
Recent images of slices of a mouse brain require ...