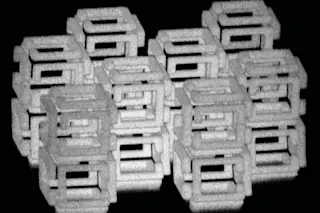
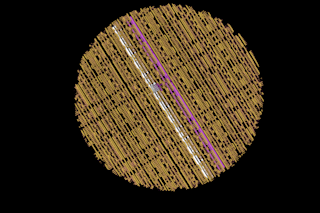
Hanging from the ceiling of Ned Seeman’s office, suspended by fishing wire, is a model of . . . um, something. Its half dozen coils of plastic tubing are paired up in a way that resembles the double helix of a DNA molecule, but DNA was never like this. Instead of forming one simple, linear strand, the plastic coils of this thing snake around, separating and coming together again to trace out a complex, cube-shaped framework. Across the room, on a table between two windows, sits an intricate assembly of sticks and balls. It’s the sort of thing a young boy might make with a set of Tinkertoys--assuming the set was large enough and the boy patient enough--but in the office of a chemistry professor at New York University, it is somewhat disconcertingly playful.
After the office, Seeman’s laboratory is a bit of a disappointment--it holds nothing you can’t see ...