Winners
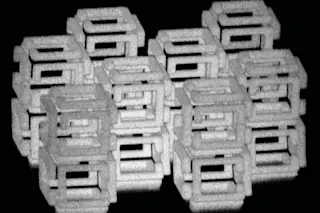
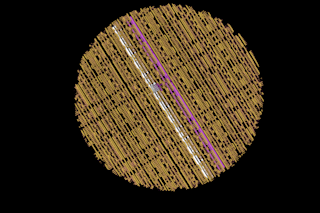
Buckyballs All in a Row
IBM’s Molecular Abacus
INNOVATOR: James Gimzewski
Ever since scientists discovered the carbon 60 molecule--a structure shaped like two geodesic domes and also known as a Buckminsterfullerene--back in the mid-1980s, they have been struggling to find something practical to do with it. James Gimzewski, a physicist at ibm’s Zurich Research Laboratory in Switzerland, has now succeeded--sort of.
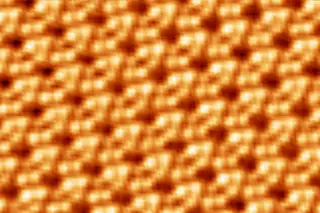
It all started when Gimzewski and his colleagues at ibm began playing around with a scanning-tunneling microscope, or stm, which has probes so microscopically thin that it can glide over a surface and record the presence of atoms like a blind person reading braille. When they learned that the tip can also be made to scrape up globs of atoms like a plow and leave behind microscopic furrows, they quickly began pushing around individual atoms for the fun of it, even going so far as to spell out ...